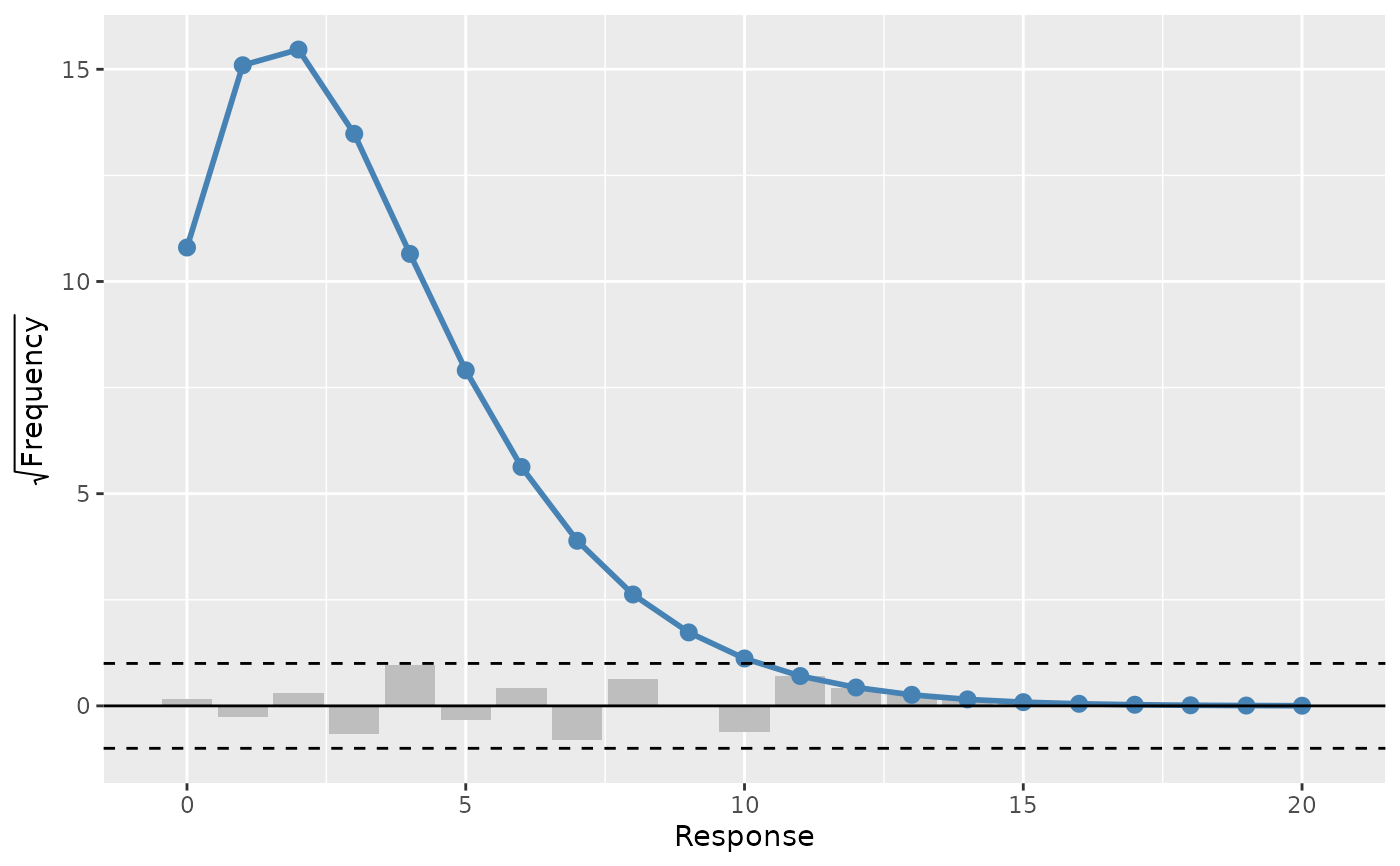
A rootogram is a model diagnostic tool that assesses the goodness of fit of
a statistical model. The observed values of the response are compared with
those expected from the fitted model. For discrete, count responses, the
frequency of each count (0, 1, 2, etc) in the observed data and expected
from the conditional distribution of the response implied by the model are
compared. For continuous variables, the observed and expected frequencies
are obtained by grouping the data into bins. The rootogram is drawn using
ggplot2::ggplot() graphics. The design closely follows Kleiber & Zeileis
(2016).
Arguments
- object
and R object to plot.
- type
character; the type of rootogram to draw.
- sqrt
logical; show the observed and fitted frequencies
- ref_line
logical; draw a reference line at zero?
- warn_limits
logical; draw Tukey's warning limit lines at +/- 1?
- fitted_colour, bar_colour, bar_fill, ref_line_colour, warn_line_colour
colours used to draw the respective element of the rootogram.
- xlab, ylab
character; labels for the x and y axis of the rootogram. May be missing (
NULL), in which case suitable labels will be used. '- ...
arguments passed to other methods.
References
Kleiber, C., Zeileis, A., (2016) Visualizing Count Data Regressions Using Rootograms. Am. Stat. 70, 296–303. doi:10.1080/00031305.2016.1173590
See also
rootogram() to compute the data for the rootogram.
Examples
load_mgcv()
df <- data_sim("eg1", n = 1000, dist = "poisson", scale = 0.1, seed = 6)
# A poisson example
m <- gam(y ~ s(x0, bs = "cr") + s(x1, bs = "cr") + s(x2, bs = "cr") +
s(x3, bs = "cr"), family = poisson(), data = df, method = "REML")
rg <- rootogram(m)
# plot the rootogram
draw(rg)
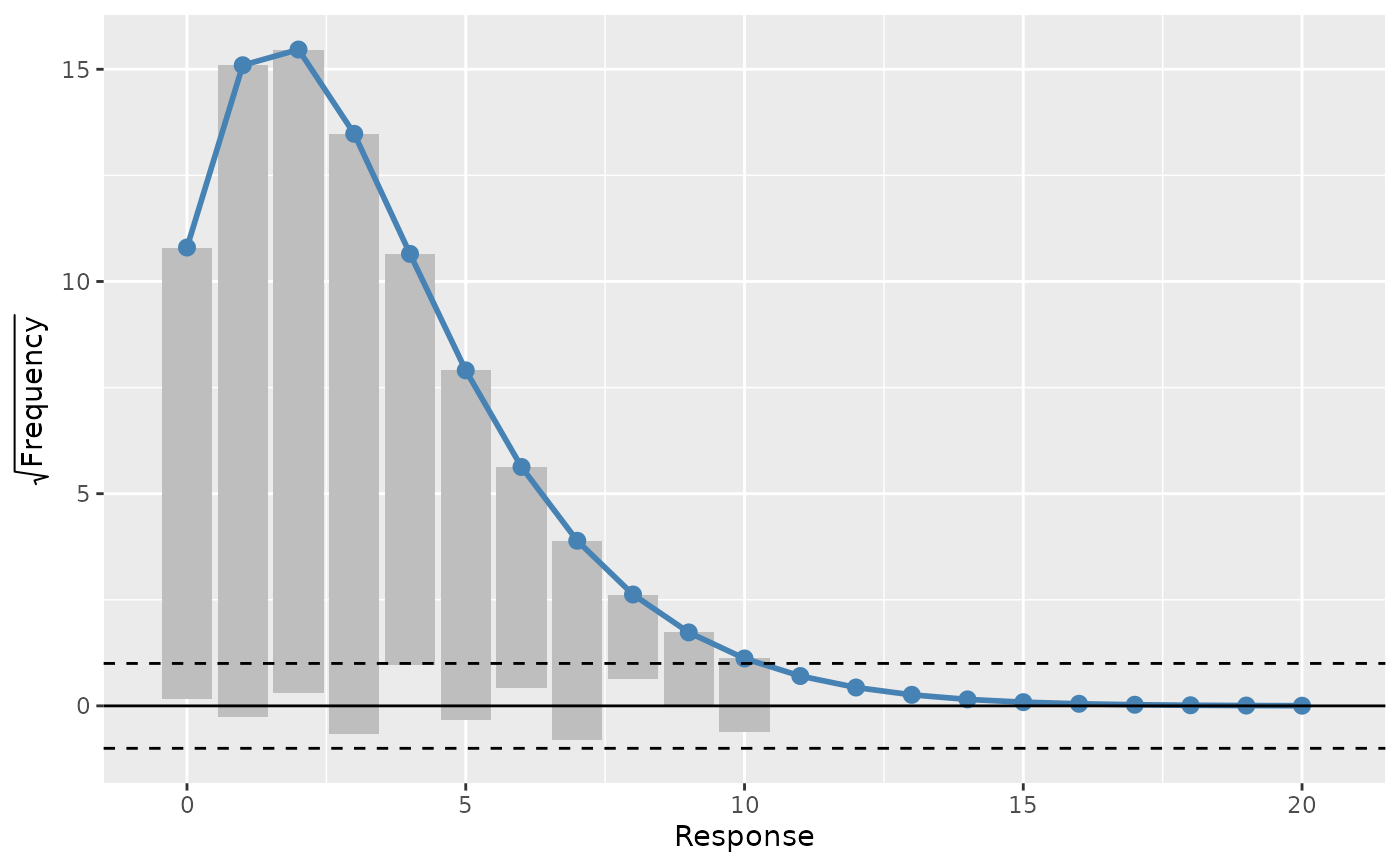 # change the type of rootogram
draw(rg, type = "suspended")
# change the type of rootogram
draw(rg, type = "suspended")