Fortify extrapolated species richness of a species pool
Source:R/fortify.poolaccum.R
fortify.poolaccum.RdPrepares a fortified version of results from
vegan::poolaccum() objects.
Arguments
- model, x
an object of class
vegan::poolaccum().- data
original data set. Currently ignored.
- alpha
level of quantiles for envelopes shown (default 0.05).
- ...
additional arguments passed to
vegan::summary.poolaccum(), notablydisplayto control which indices should be computed.
Value
A data frame with columns index, size, richness, lower,
upper, and std_dev, containing the richness index, permutation richness
estimator, sample size, upper and lower \(1 - \alpha\)
quantile interval, and standard deviation of permutation estimates,
respectively.
Examples
library("vegan")
library("ggplot2")
data(BCI)
pool <- poolaccum(BCI)
df <- fortify(pool)
df
#> # A tibble: 240 × 6
#> index size richness lower upper std_dev
#> <fct> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 S 3 139. 125. 150. 6.78
#> 2 S 4 151. 140. 161. 5.43
#> 3 S 5 159. 147 171. 5.74
#> 4 S 6 165. 155. 176 5.26
#> 5 S 7 170. 162. 180. 4.84
#> 6 S 8 175. 166. 185. 5.32
#> 7 S 9 178. 170 188. 5.27
#> 8 S 10 181. 172. 193. 5.59
#> 9 S 11 184. 175. 196. 5.34
#> 10 S 12 187. 177. 198. 5.29
#> # ℹ 230 more rows
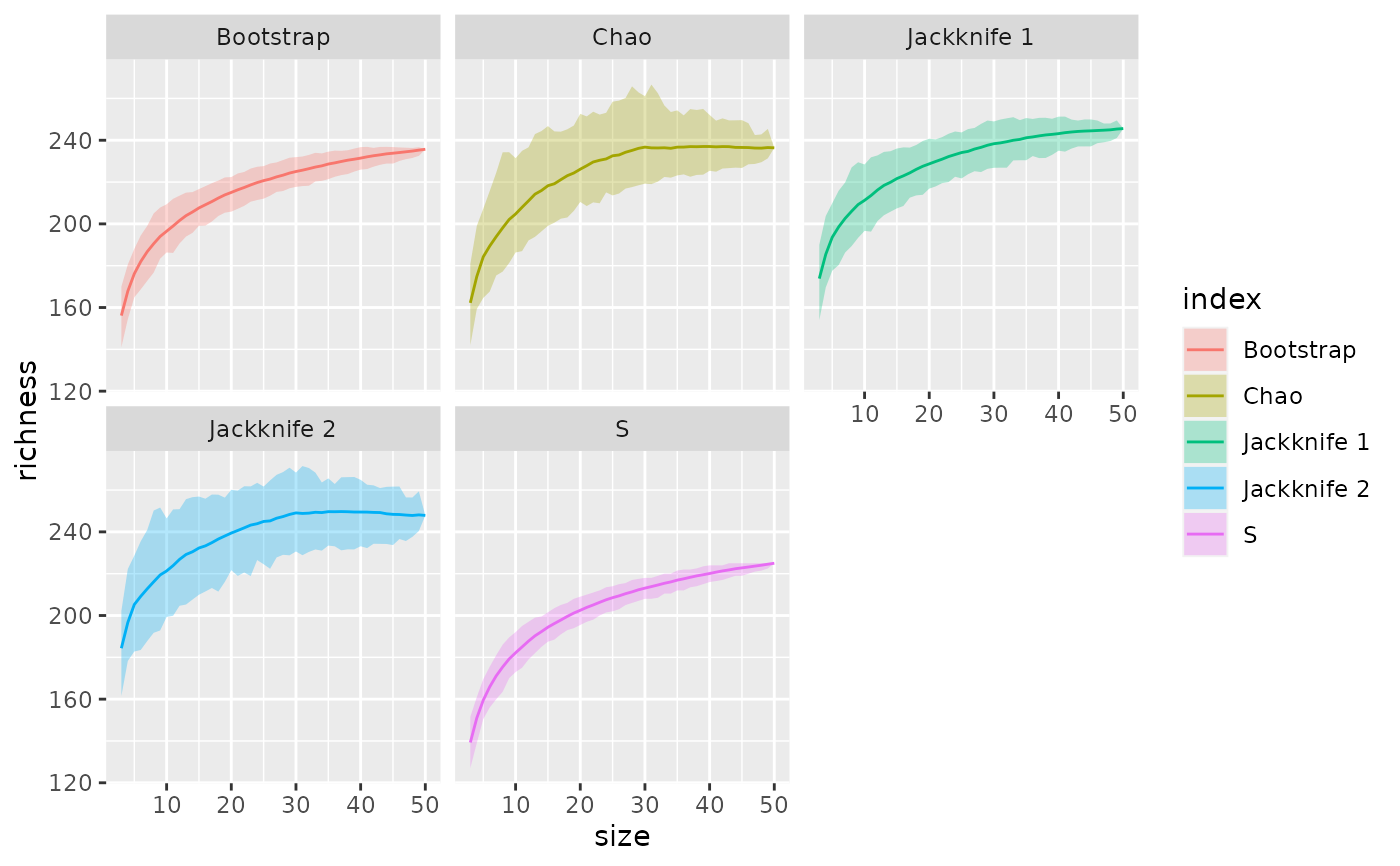
ggplot(df, aes(x = size, y = richness, colour = index)) +
geom_ribbon(aes(ymin = lower, ymax = upper, x = size, fill = index),
alpha = 0.3, inherit.aes = FALSE) +
geom_line() +
facet_wrap(~ index)